We seek to understand the genetic basis of common diseases and to translate genetic association data into biological insights. Most of our research involves the development of new statistical methods and the application of these methods to large-scale genetic datasets.
Genetic architecture of common and rare variation 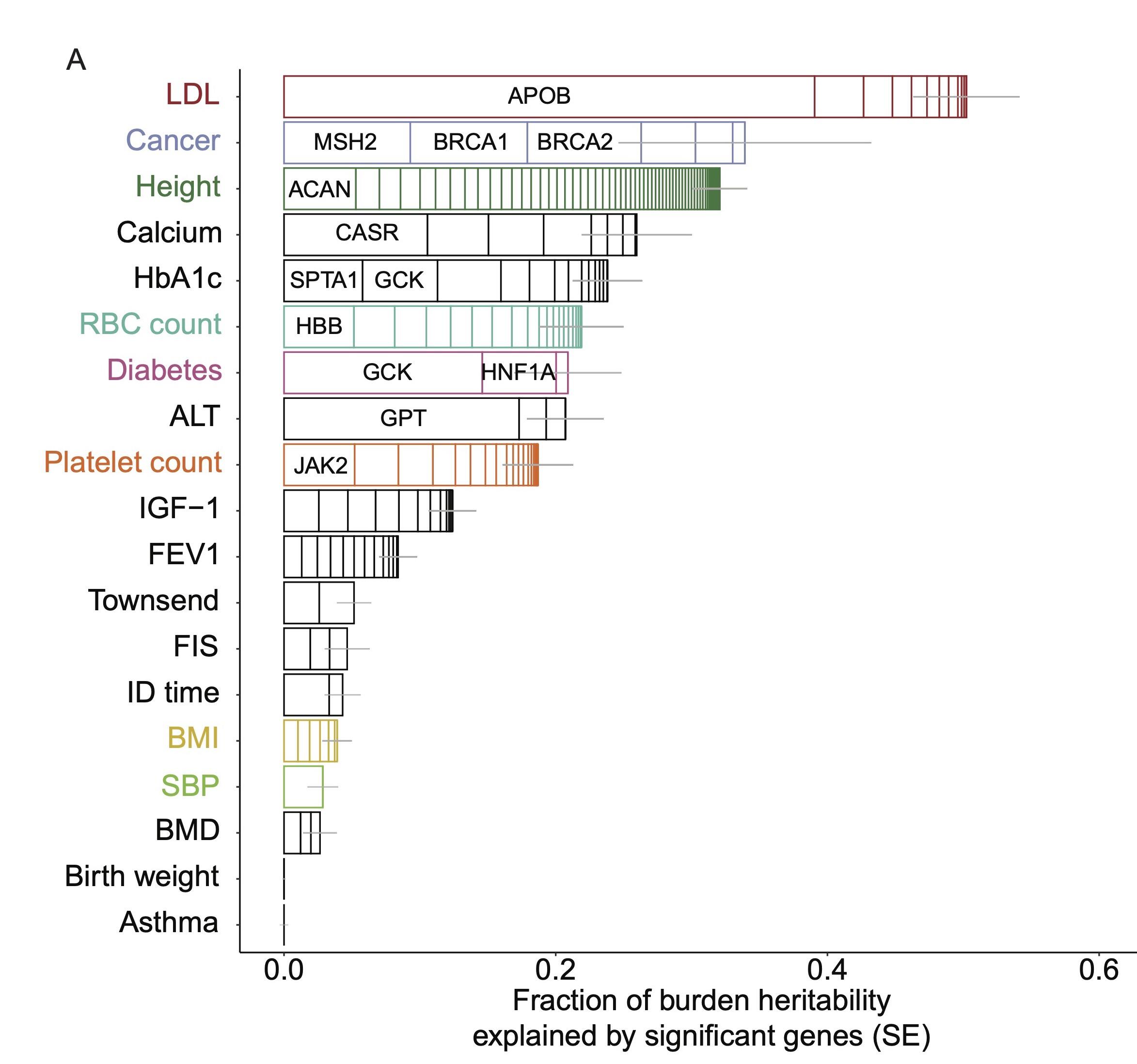
Both common and rare genetic variants influence common diseases and complex traits. We have developed methods to characterize the genetic architecture of common and rare variants, with a particular focus on how and why they differ. A major focus of our ongoing research is the development of methods to analyze rare variant association data.
Weiner*, Nadig* et al. Nature
Weiner et al. 2022 AJHG
O’Connor 2021 Nat Genet
O’Connor et al. 2019 AJHG
Linkage disequilibrium graphical models 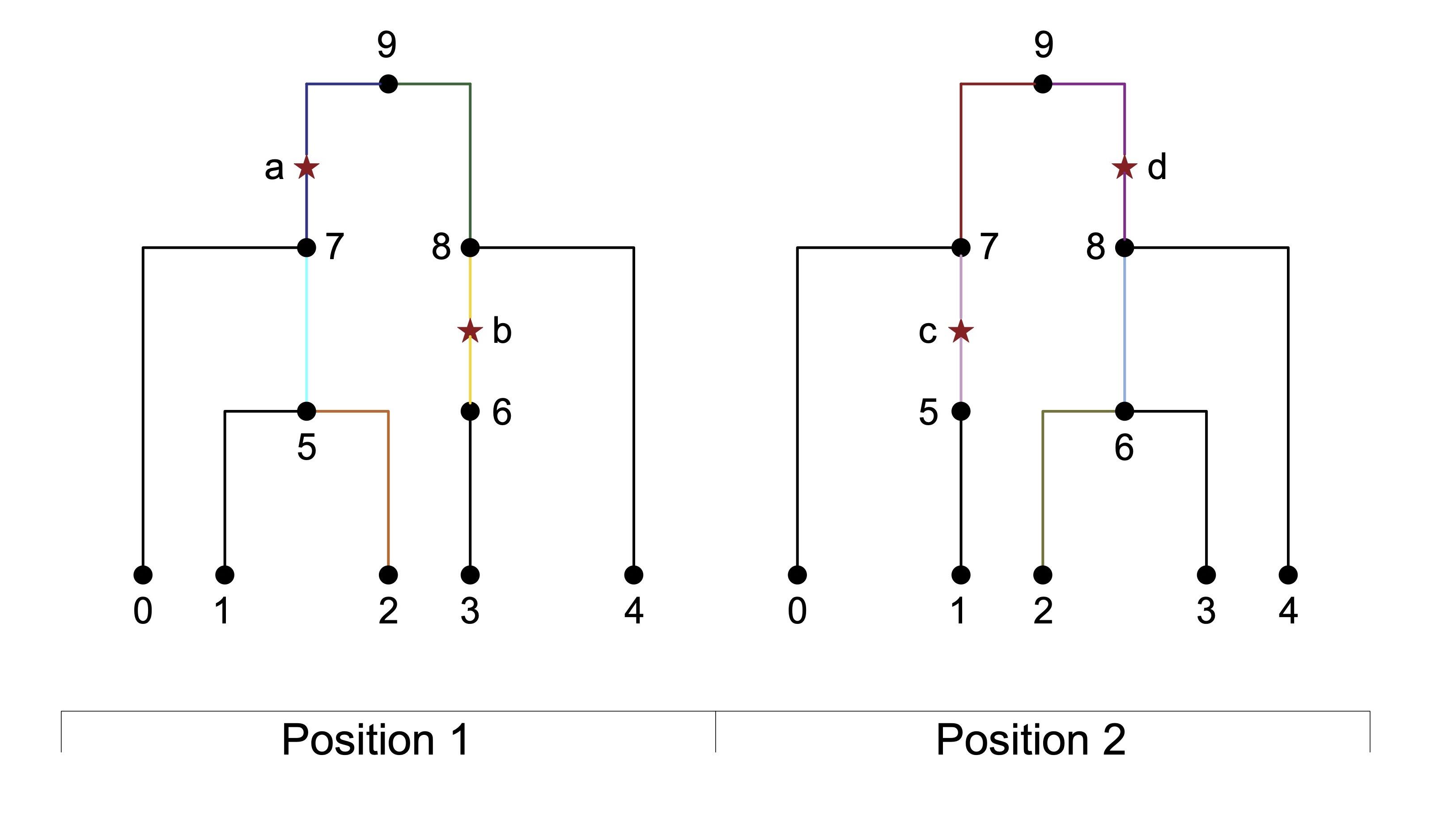
Statistical methods to analyze common variants must account for linkage disequilibrium (LD), especially for studies that are ancestrally diverse. We developed an approach to model LD using extremely sparse LD graphical models (LDGMs), which we infer from genome-wide genealogies. We are developing applications of LDGMs to polygenic risk prediction and heritability partitioning.
Salehi*, Wohns* et al. 2023 Nat Genet
Pleiotropy and causality 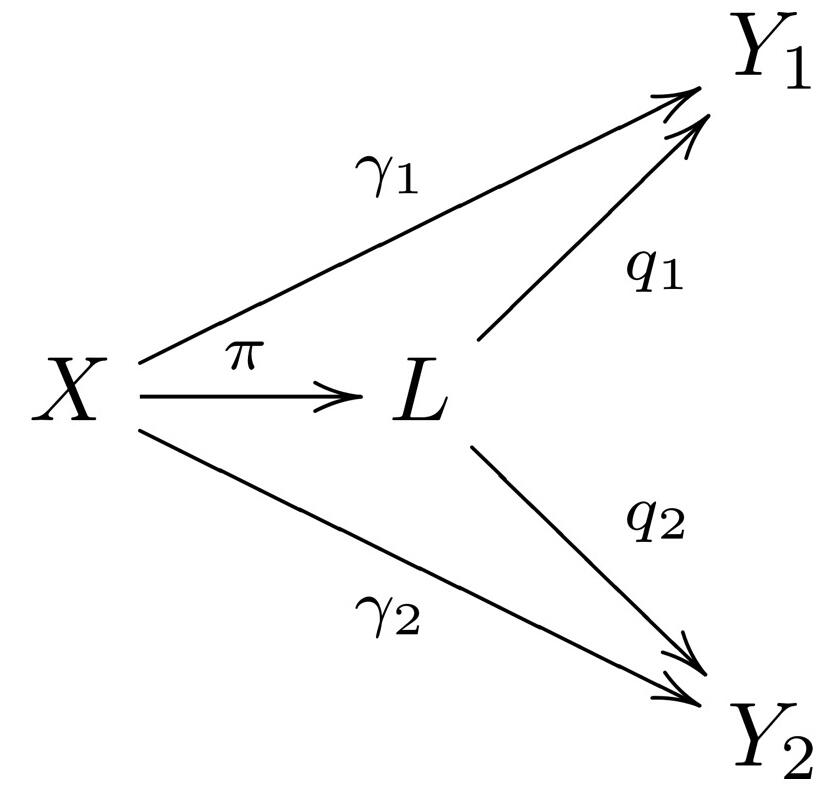
Most disease-associated SNPs are pleiotropically associated with other traits as well, pointing to shared mechanisms or even causal relationships. We have developed methods to characterize genetic and biological relationships among genetically correlated traits.
Ballard and O’Connor 2022 AJHG
O’Connor and Price 2018 Nat Genet